1: A quick use case for MICADO at the ELT¶
A brief introduction into using ScopeSim to observe a cluster in the LMC¶
This is a step-by-step guide. The complete script can be found at the bottom of this page/notebook.
First set up all relevant imports:
[1]:
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
%matplotlib inline
import scopesim as sim
import scopesim_templates as sim_tp
updating/loading 'index.yml'
Downloading https://scopesim.univie.ac.at/spextra/database/index.yml [Done]
updating/loading 'default_filters.yml'
Downloading https://scopesim.univie.ac.at/spextra/database/default_filters.yml [Done]
updating/loading 'default_spectra.yml'
Downloading https://scopesim.univie.ac.at/spextra/database/default_spectra.yml [Done]
updating/loading 'default_curves.yml'
Downloading https://scopesim.univie.ac.at/spextra/database/default_curves.yml [Done]
Scopesim works by using so-called instrument packages, which have to be downloaded separately. For normal use, you would set the package directory (a local folder path, local_package_folder in this example), download the required packages once, and then remove the download command.
[2]:
local_package_folder = "./inst_pkgs"
However, to be able to run this example on the Readthedocs page, we need to include a temporary directory.
Do not copy and run this code locally, it is only needed to set things up for Readthedocs!
[3]:
from tempfile import TemporaryDirectory
local_package_folder = TemporaryDirectory().name
Download the required instrument packages for an observation with MICADO at the ELT.
Again, you would only need to do this once, not every time you run the rest of the script, assuming you set a (permanent) instrument package folder.
[4]:
sim.rc.__config__["!SIM.file.local_packages_path"] = local_package_folder
sim.download_packages(["Armazones", "ELT", "MORFEO", "MICADO"])
astar.scopesim.server.database - Gathering information from server ...
astar.scopesim.server.database - Connection successful, starting download ...
Downloading Armazones: 100%|██████████| 74.3k/74.3k [00:00<00:00, 632kB/s]
Extracting Armazones: 100%|██████████| 10/10 [00:00<00:00, 3139.68it/s]
Downloading ELT : 100%|██████████| 53.4k/53.4k [00:00<00:00, 25.7MB/s]
Extracting ELT : 100%|██████████| 16/16 [00:00<00:00, 5517.91it/s]
Downloading MORFEO : 100%|██████████| 1.38M/1.38M [00:02<00:00, 562kB/s]
Extracting MORFEO : 100%|██████████| 12/12 [00:00<00:00, 912.45it/s]
Downloading MICADO : 100%|██████████| 14.4M/14.4M [00:27<00:00, 549kB/s]
Extracting MICADO : 100%|██████████| 121/121 [00:00<00:00, 846.82it/s]
[4]:
[PosixPath('/tmp/tmpgjh6i23k/Armazones.zip'),
PosixPath('/tmp/tmpgjh6i23k/ELT.zip'),
PosixPath('/tmp/tmpgjh6i23k/MORFEO.zip'),
PosixPath('/tmp/tmpgjh6i23k/MICADO.zip')]
Now, create a star cluster using the scopesim_templates package. You can ignore the output that is sometimes printed. The seed argument is used to control the random number generation that creates the stars in the cluster. If this number is kept the same, the output will be consistent with each run, otherwise the position and brightness of the stars is randomised every time.
[5]:
cluster = sim_tp.stellar.clusters.cluster(mass=1000, # Msun
distance=50000, # parsec
core_radius=0.3, # parsec
seed=9002)
imf - sample_imf: Setting maximum allowed mass to 1000
imf - sample_imf: Loop 0 added 1.26e+03 Msun to previous total of 0.00e+00 Msun
updating/loading 'filter_systems/etc/index.yml'
Downloading https://scopesim.univie.ac.at/spextra/database/filter_systems/etc/index.yml [Done]
updating/loading 'filter_systems/etc/V.dat'
Downloading https://scopesim.univie.ac.at/spextra/database/filter_systems/etc/V.dat [Done]
updating/loading 'libraries/ref/index.yml'
Downloading https://scopesim.univie.ac.at/spextra/database/libraries/ref/index.yml [Done]
updating/loading 'libraries/ref/vega.fits'
Downloading https://scopesim.univie.ac.at/spextra/database/libraries/ref/vega.fits [Done]
Next, make the MICADO optical system model with OpticalTrain. Observe the cluster Source object with the .observe() method and read out the MICADO detectors with .readout(). This may take a few moments on slower machines.
The resulting FITS file can either be returned as an astropy.fits.HDUList object, or saved to disk using the optional filename parameter
[6]:
micado = sim.OpticalTrain("MICADO")
micado.observe(cluster)
hdus = micado.readout()
# micado.readout(filename="TEST.fits")
FOVs: 0%| | 0/1 [00:00<?, ?it/s]
FOV effects: 0%| | 0/2 [00:00<?, ?it/s]
FOV effects: 50%|█████ | 1/2 [00:00<00:00, 6.24it/s]
FOV effects: 100%|██████████| 2/2 [00:00<00:00, 7.17it/s]
FOVs: 100%|██████████| 1/1 [00:00<00:00, 1.89it/s]
Image Plane effects: 100%|██████████| 1/1 [00:00<00:00, 5295.84it/s]
astar.scopesim.detector.detector_array - Extracting from 1 detectors...
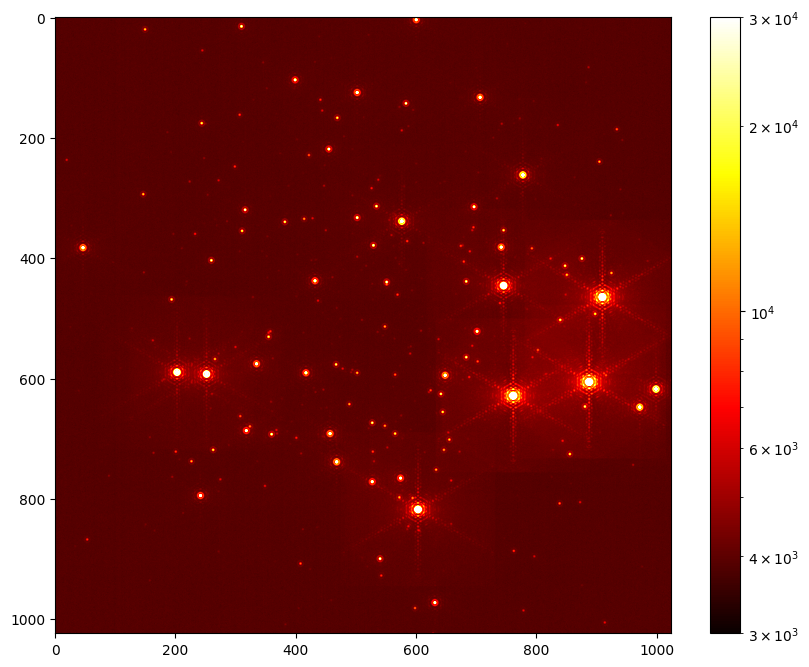
Display the contents the first HDU
[7]:
plt.figure(figsize=(10,8))
plt.imshow(hdus[0][1].data, norm=LogNorm(vmax=3E4, vmin=3E3), cmap="hot")
plt.colorbar()
[7]:
<matplotlib.colorbar.Colorbar at 0x7f758f25e990>
Complete script¶
Included below is the complete script for convenience, including the downloads, but not including the plotting.
[8]:
import scopesim as sim
import scopesim_templates as sim_tp
#sim.download_packages(["Armazones", "ELT", "MORFEO", "MICADO"])
cluster = sim_tp.stellar.clusters.cluster(mass=1000, # Msun
distance=50000, # parsec
core_radius=0.3, # parsec
seed=9002)
micado = sim.OpticalTrain("MICADO")
micado.observe(cluster)
hdus = micado.readout()
# micado.readout(filename="TEST.fits")
imf - sample_imf: Setting maximum allowed mass to 1000
imf - sample_imf: Loop 0 added 1.26e+03 Msun to previous total of 0.00e+00 Msun
FOVs: 0%| | 0/1 [00:00<?, ?it/s]
FOV effects: 0%| | 0/2 [00:00<?, ?it/s]
FOV effects: 50%|█████ | 1/2 [00:00<00:00, 6.59it/s]
FOV effects: 100%|██████████| 2/2 [00:00<00:00, 7.38it/s]
FOVs: 100%|██████████| 1/1 [00:00<00:00, 1.95it/s]
Image Plane effects: 100%|██████████| 1/1 [00:00<00:00, 5059.47it/s]
astar.scopesim.detector.detector_array - Extracting from 1 detectors...
[ ]: